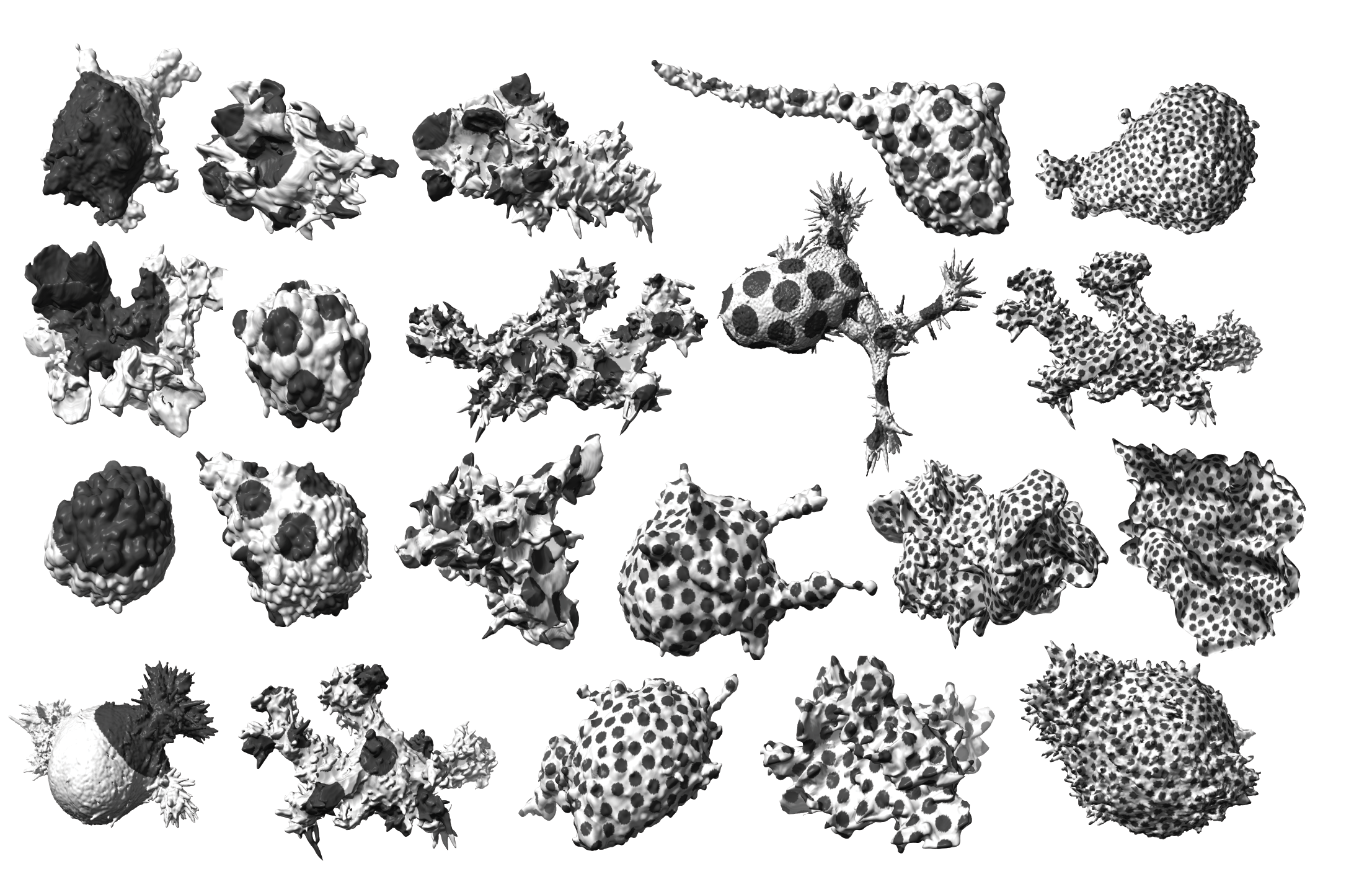
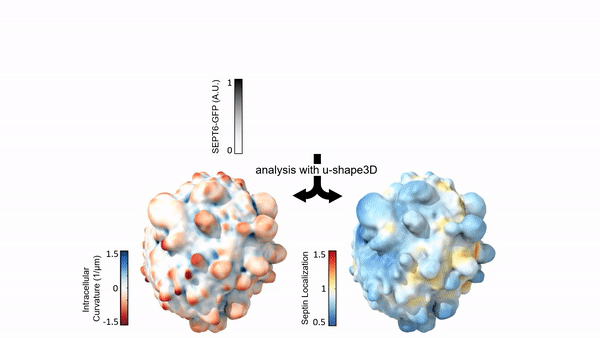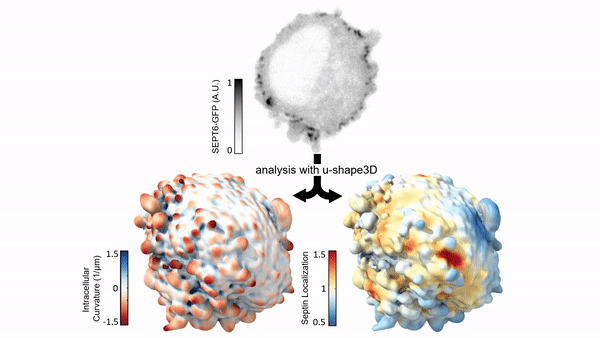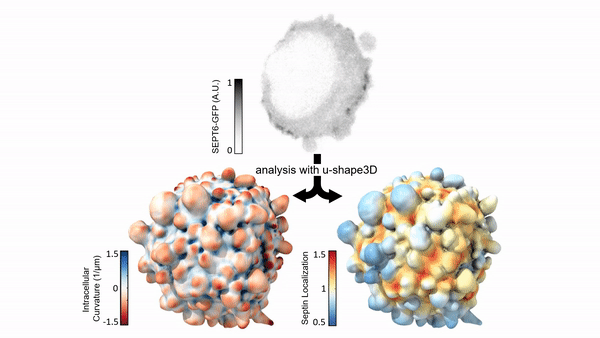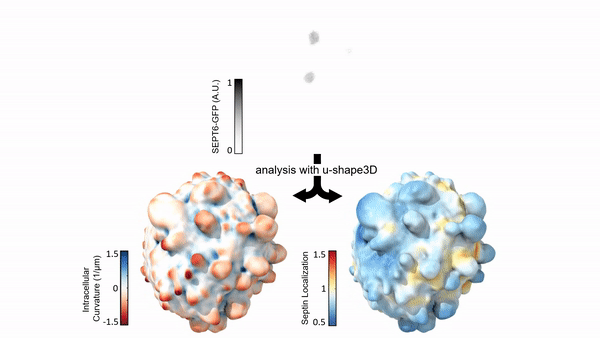
Measuring & Modeling Non-Genetic Mechanisms of Cancer Metastasis & Drug Resistance
Underlying cancer metastasis and drug resistance is an ability of cells to cope with ever-changing and often growth-challenging environmental conditions. Some of this adaptive power roots in clonal cell selection, but it also occurs via acute cell functional plasticity. Our lab collaborates with microscopy developers, chemists, developmental biologists, and clinicians to take an ever-closer look at the multi-factorial molecular processes underlying cellular plasticity. We develop cutting-edge computer vision approaches and statistical methods for causal inference to identify integrated circuits of molecular signaling, cell architecture, and cell shape that control the cellular decisions behind plastic behaviors.