Our Research
The broad research interest of Fei Wang lab is in dissecting molecular mechanisms of essential cellular events in eukaryotic cell development. Currently, my laboratory investigates how cells preserve the fidelity of gametogenesis under environmental and physiological stress. Gametogenesis, the process by which diploid precursors undergo meiosis and differentiation to produce haploid gametes, is remarkably sensitive to temperature and other stressors. Even mild perturbations tolerated by mitotically dividing cells can disrupt meiosis, leading to infertility or defective gametes. This heightened sensitivity reflects the competing demands faced by germ cells: they must complete complex developmental transitions while responding flexibly to environmental challenges. The mechanisms that safeguard meiotic integrity under such conditions have long remained poorly understood. In somatic cells, two conserved pathways—stress granules (SGs) and autophagy—serve as major quality-control systems under stress. SGs sequester untranslated mRNAs and RNA-binding proteins to pause translation and preserve key transcripts, while autophagy recycles damaged or unneeded components through autophagosome-mediated degradation, restoring proteostasis and energy balance. How these systems cooperate during meiosis, when transcriptional and translational reprogramming, organelle remodeling, and proteome renewal occur simultaneously, has remained largely unexplored.
Using Saccharomyces cerevisiae (budding yeast) as a powerful model, my group integrates genetics, biochemistry, live-cell imaging, proteomics, and computational approaches to uncover how autophagy and RNA regulation ensure gamete quality under stress. Supported by Welch and NIGMS(R01 and R35) fundings and an endowed scholarship to Dr. Fei Wang (Nancy Cain and Jeffrey A. Marcus Scholar in Medical Research, in Honor of Dr. Bill S. Vowell), our research has defined a unified quality-control network that integrates autophagy, RNA regulation, and stress response to preserve reproductive fidelity.
Impact of Our Research
Gametogenesis is one of the most stress-sensitive developmental processes in eukaryotes, from yeast to humans. Understanding how stress-responsive mechanisms safeguard gametogenesis has broad implications for fertility biology, environmental adaptation, and reproductive health. Our re will also provide foundational knowledge that could inform future investigations into protein misfolding disorders, infertility syndromes, and stress adaptation in agricultural systems.
Autophagy is a lysosomal degradation pathway. Numerous human diseases, such as cancer, and infectious and developmental diseases have been linked to aberrant autophagy. Besides its disease relevance, autophagy is a unique system for investigating the basic principles of organellar biogenesis because, unlike many organelles that are rich in membrane and protein components (endoplasmic reticulum, mitochondria, etc), autophagosomes are relatively simple. Autophagosomal biogenesis responds to a variety of internal and external signals, making fine genetic or chemical manipulation possible for understanding molecular mechanisms and defining new targets for disease treatment. Canonically, nascent autophagosomes fuse with lysosomes to deliver their cargos for degradation. However, under certain circumstances, autophagosomes appear to also have degradation-independent functions, for example, protein secretion. We propose that the canonical and non-canonical roles of autophagosomes co-operate to guide cells through critical stages of development, such as meiosis, when the cell is programmed to undergo drastic cellular content degradation and structural remodeling. We demonstrated that, while autophagy is dispensable for mitosis, it is essential for meiosis at multiple stages (Dev Cell, 2020). We discovered that the conserved phosphatase Cdc14 directly activates autophagy during meiotic divisions, establishing the first mechanistic, meiosis-specific “on-switch” for autophagy (J. Cell Biol., 2022), and that the meiosis-specific RNA-binding protein Rim4—an essential repressor of post-transcriptional meiotic gene expression—is both a selective autophagy substrate and the master regulator of meiotic stress granules (Cell Reports, 2023a,b; Nat. Commun., 2025). These findings reveal how the evolutionarily conserved enzymes PKA, Cdc14, and 14-3-3 proteins coordinate Rim4’s phosphorylation state and spatiotemporal dynamics to couple translational repression, autophagic degradation, and stress adaptation with meiotic progression. Our ongoing studies will keep shedding light directly on autophagy functions in eukaryotic gametogenesis, the production of sperm (spermatogenesis) and oocytes (oogenesis).
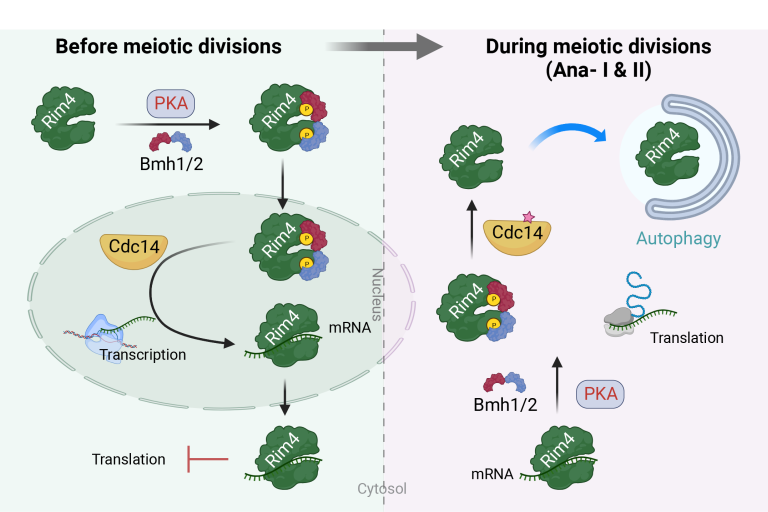 Regulation of Rim4 Distribution, Function, and Stability during Meiosis by PKA, Cdc14, and 14-3-3 Proteins
Regulation of Rim4 Distribution, Function, and Stability during Meiosis by PKA, Cdc14, and 14-3-3 Proteins The extraordinary thermosensitivity of gametogenesis in many species had been observed for decades, but its molecular origin was unknown. We discovered that meiotic cells form stress granules (SGs) at temperatures nearly 10 °C lower than those triggering SGs in mitotic cells. This hypersensitivity stems from Rim4, whose third RNA-recognition motif seeds SG formation upon mild heat stress, regulated by site-specific phosphorylation. These “Rim4-seeded SGs” act as reversible protective hubs that transiently pause meiosis to safeguard its fidelity.
We demonstrated that Rim4-seeded stress granules connect temperature sensing to meiotic control. (1) This was the first demonstration that SGs play a role in yeast sexual reproduction. It provides a molecular explanation for the long-oberved extreme thermosensitivity of gametogenesis—exemplified by the external location of testes, which maintains spermatogenesis at temperatures 2–7 °C below core body temperature. (2) We demonstrated that Rim4 is the master regulator of meiotic SGs, functionally analogous to G3BP1, a central RNA-binding protein that nucleates stress granule assembly in mammalian cells. Our finding bridges yeast and metazoan stress-response mechanisms. (3) We established a novel "Rim4 seed" model that directly connects environmental sensing to meiotic cell cycle control via SG dynamics, proposing a mechanism that could underlie temperature-dependent fertility in diverse eukaryotes. (4) By engineering Rim4, we induced SG formation in the absence of heat, demonstrating that SG assembly alone can halt meiosis and hinting at possible strategies to modulate gamete quality through controlled SG manipulation.
Proteins arising from small ORFs that are substantially smaller than ~50 residues are considered small proteins (SPs) which historically tend to be ignored. Recent studies, however, suggest that hundreds or even thousands of SPs were synthesized in bacteria and eukaryotes with functions implicated in diverse processes such as spore formation, cell division, movement of molecules across the membrane, enzymatic activity, and signal transduction. SPs are small and prone to interact with their targets, often on the membrane, to modulate their activities, therefore with great potential in clinical application. We are interested in elucidation of a molecular mechanism related to SPs: the targeting machinery, their structure nature, interactions with other molecules as well as their turnover in eukaryotes.
Yeast genetics; Live cell imaging; Electronic microscopy; Proteomic analyses of protein interactomes; Protein biochemistry; Biochemical reconstitution of purified proteins and lipids; in vivo protein labeling
 FM
FM  Lab
Lab Meet the Principal Investigator

Fei Wang, Ph.D.
Fei Wang received his B.S. in Genetics and M.S. in Gene Therapy from Fudan University, China, in 1997 and 2001, respectively. He attended graduate school at UMass-Amherst and earned a Ph.D. in Biochemistry and Molecular Biology with Danny Schnell, Ph.D. He was then trained as a postdoctoral fellow in the research group of Vlad Denic, Ph.D., at Harvard University on defining the molecular mechanism of post-translational translocation of the single pass integral ER membrane proteins: Tail-anchored proteins.
In 2017, Dr. Wang joined the faculty at UT Southwestern in the Center for Autophagy Research, Department of Internal Medicine with a secondary appointment in the Department of Cell Biology.
Meet the Lab Members
Rudian Zhang, Ph.D.
Research Scientist
Research Interests:
Autophagic degradation of protein aggregation/meiotic translational control

Jin Li, Ph.D.
Postdoctoral Researcher
Research Interests: The relationship between SGs (stress granules) and cellular organelles during the progress of meiosis

Interested in joining?
- Dr. Fei Wang
- Department of Cell Biology
- UT Southwestern Medical Center, NL6. 120F 6000 Harry Hines Blvd. Dallas, TX 75390
- 2146488491
Lab Alumni
Jayaprakash Monala, Ph.D.
Postdoctoral Researcher

Akshay Chellappa
Research Technician

Wenzhi Feng (Postdoc)
Jiajia Li (Research Assistant)
Suhong Qian (Research Associate)
Ella Zhang (STARs program student)
Joel Maliyil (STARs program student)
Featured Publications
Rim4-seeded stress granules connect temperature sensing to meiotic regulation.
Zhang R, Li S, Feng W, Qian S, Chellappa AJ, Wang F, 2025 Jul Nat Commun 1 16 5566Autophagy-Mediated Surveillance of Rim4-mRNA Interaction Safeguards Programmed Meiotic Translation
Rudian Zhang, Wenzhi Feng, Suhong Qian, Fei Wang 2023 Cell Reports DOI: 10.1016/j.celrep.2023.113051Cdc14 spatiotemporally dephosphorylates Atg13 to activate autophagy during meiotic divisions.
Feng W, Argüello-Miranda O, Qian S, Wang F, 2022 May J Cell Biol 5 221Autophagy of an Amyloid-like Translational Repressor Regulates Meiotic Exit.
Wang F, Zhang R, Feng W, Tsuchiya D, Ballew O, Li J, Denic V, Lacefield S, 2020 Jan Dev. Cell 2 52 141-151.e5The Get1/2 transmembrane complex is an endoplasmic-reticulum membrane protein insertase.
Wang F, Chan C, Weir NR, Denic V 2014 Aug Nature 7515 512 441-4The mechanism of tail-anchored protein insertion into the ER membrane.
Wang F, Whynot A, Tung M, Denic V 2011 Sep Mol. Cell 5 43 738-50Structural basis for tail-anchored membrane protein biogenesis by the Get3-receptor complex.
Stefer S, Reitz S, Wang F, Wild K, Pang YY, Schwarz D, Bomke J, Hein C, Löhr F, Bernhard F, Denic V, Dötsch V, Sinning I 2011 Aug Science 6043 333 758-62Contact Us
Fei Wang, Ph.D.
Assistant Professor
Department of Cell Biology and Biochemistry
Nancy Cain and Jeffrey A. Marcus Scholar in Medical Research, in Honor of Dr. Bill S. Vowell
Email Dr. Wang
Phone: 214-648-8491
UT Southwestern Medical Center, NL6. 120F
6000 Harry Hines Blvd.
Dallas, TX 75390
Join Our Lab
We are committed to providing a pleasant environment to energetic and motivated researchers interested in science who like to work independently and on a small team. To request a position as a postdoctoral fellow, graduate student, or technician, please contact us.
