Gene expression is a highly dynamic process. To regulate RNA synthesis and degradation, cells have a variety of tools for regulating gene expression.
However, many childhood cancers are driven by the loss of these regulatory tools. For example, the loss of one of these tools, known as microRNAs, can lead to a variety of rare childhood tumors. Many other cancers bear mutations in transcription factors, the proteins that give a cell its identity by determining which genes are expressed. Yet other cancers are driven by mutations in another regulatory tool, the chromatin remodeling complexes.
We study how dysregulation of RNA synthesis and degradation drives childhood cancers with the ultimate goal of identifying new therapeutic vulnerabilities to exploit in treating them.
Wilms tumor
Wilms tumor is the most common pediatric kidney cancer. Many cases are driven by mutations in transcription factors, chromatin remodeling factors, or microRNA processing genes. We are studying how these mutations affect gene expression and drive tumor formation.
These studies are performed both in vitro using cell-line models and in vivo by generating genetically engineered mouse models. In addition, we continue to an Wilms tumors to fully describe the landscape of gene dysregulation mechanisms.

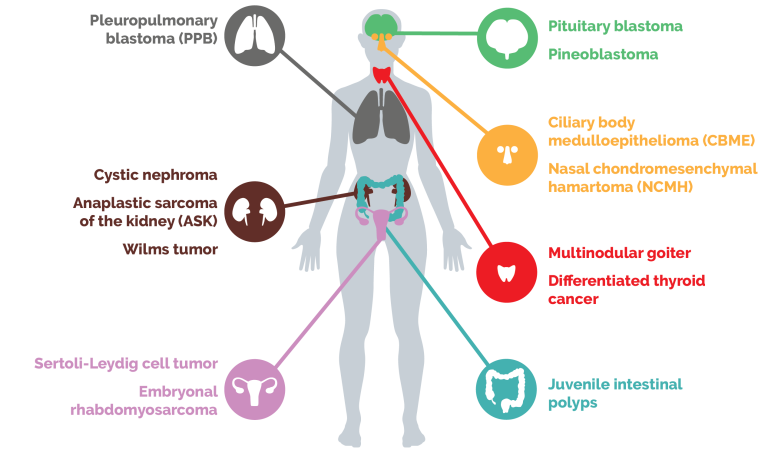
DICER1-related cancers
Mutations in the microRNA-processing gene DICER1 predispose individuals to a spectrum of rare pediatric cancers.
We use a combination of genomic analysis, cell-based models, and in vivo systems to understand how DICER1 dysfunction alters gene expression and drives tumorigenesis, in terms of mechanisms of microRNA loss, tumor-specific vulnerabilities, and translational opportunities.
Pineoblastoma
While other cancer types may be driven by partial loss of microRNAs, the pediatric brain tumor pineoblastoma is unique because it can form with complete loss of microRNAs. To understand what makes these tumors function, we built a genetic mouse model of pineoblastoma by knocking out microRNA factors. Interestingly, these tumors harness some of the same tumor-driving pathways we see in DROSHA-driven Wilms tumor. We are continuing to use this model to identify new ways to block these tumor-driving pathways.