The Past (mostly)
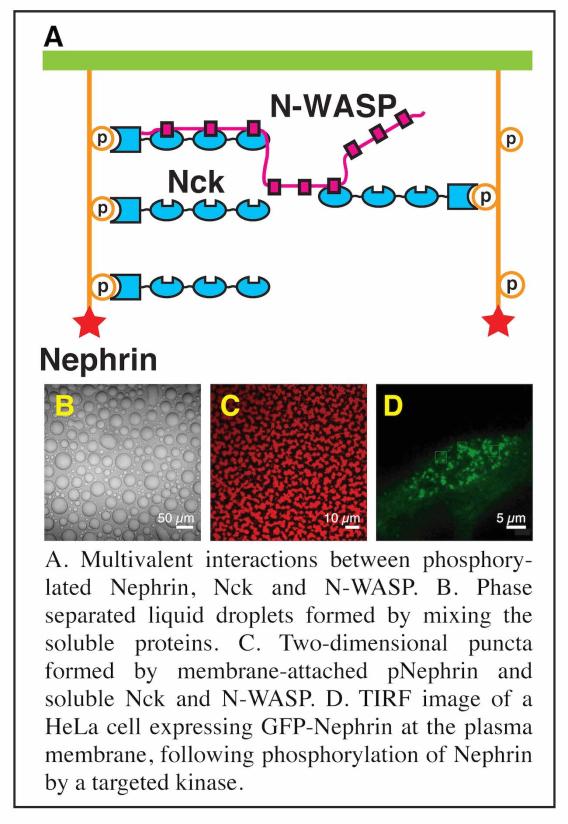
Prior to ~2010, research in our lab focused on understanding molecular scale mechanisms underlying actin regulatory signaling pathways, primarily using NMR spectroscopy and x-ray crystallography. We continue to do some work in this area. For example, we are working to understand the µs-ms conformational dynamics of the actin monomer in different nucleotide states, to learn how nucleotide controls assembly of actin filaments. But most of our recent efforts have shifted to studies of biomolecular condensates, both three-dimensional compartments in the cytoplasm and nucleoplasm, as well as two-dimensional clusters of actin-regulatory signaling molecules at membranes. By connecting concepts in physical chemistry and polymer science with biological molecules, we have helped establish important concepts in this area. We showed that diverse multivalent molecules can undergo assembly and concomitant liquid-liquid phase separation (LLPS) in vitro and in cells, creating structures that closely resemble natural condensates.
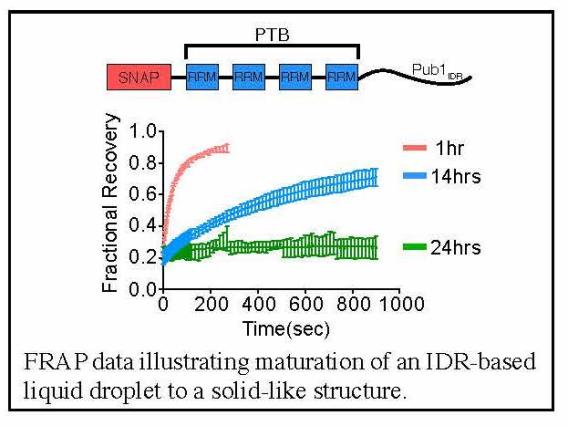
Through this idea of multivalency-driven LLPS we explained behaviors of multidomain proteins, intrinsically disordered proteins and nucleic acids. We showed that phase separated liquids formed by disordered proteins mature to solids over time through assembly into amyloid-like fibers, providing insight into regulation of RNA granules and neurodegenerative disease (in collaboration with Roy Parker’s lab).
We demonstrated that phase separation can be rapidly controlled by covalent modifications, and also by genetic changes, which both act through altering valencies of the interacting species.
Through reconstitution of a 13-component signaling pathway in vitro from purified components identified biochemical and cellular functions that uniquely arise because of the macroscopic organization of signaling clusters in T cells (in collaboration with Ron Vale’s lab).
Finally, we developed a generalizable framework to explain the regulated composition of phase separated structures.
In a broad sense, we illustrated how the complexities of cellular body formation, regulation and function can be reduced to biochemically tractable problems and simple rules.
The Future
Molecular features driving phase separation
Our future studies, described in more detail in the sections of individual lab members, are directed in three broad areas. First, we are working to understand the physical principles that enable different classes of macromolecules to undergo LLPS. This includes the surface properties and inter-domain linker sequences of multi-domain proteins involved in signal transduction, as well as the sequence properties of disordered elements such as those found in RNA binding proteins. On the latter, we are interested both in sequence features that drive LLPS, as well as those that promote maturation to amyloid-like fibers. We also are using these various systems to understand how the physical properties of molecules dictate the macroscopic properties of their phase separated structures. These studies involve low-throughput protein design followed by biochemical and cell biological analyses, and also development of high-throughput in vitro screens to examine the properties of thousands of proteins at a time.
To ground these investigations in physical theory, we are working collaboratively with several groups in condensed matter physics—Milo Lin, Rohit Pappu and Ned Wingreen.
Composition and function of condensates
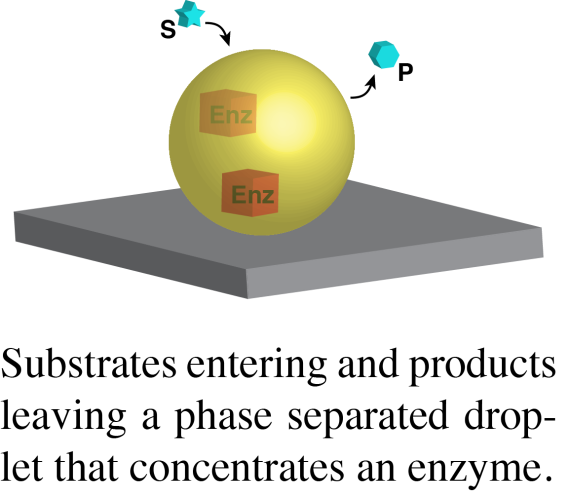
In a second area, we are working to understand the compositions, and biochemical/biological functions of natural condensates. We currently have projects involving PML nuclear bodies, yeast P bodies, chromatin domains and actin regulatory signaling clusters. But we are always looking for new systems to explore new biology and reveal general principles regarding condensate structure and function. Our goals in these projects are to first quantify the absolute concentrations of the dominant molecules in condensates in cells, and then to use this information to generate representative biochemical reconstitutions of the structures in vitro, either in solution or on lipid bilayers. With reconstitutions in hand, we seek to understand how phase separation affects the physical properties and chemistry (binding, catalysis) of the concentrated proteins. Through comparisons with parallel studies of simpler model phase separating systems, we hope to reveal general principles regarding how the molecular and macroscopic properties of phase separated structures control the molecules within them. Analogous to our published work on T cell signaling will use these reconstitutions to design precise cellular perturbations, to understand how phase separation is used to achieve biological functions.

Ultrastructure of condensates
In a final area, we are just beginning studies of the ultrastructure of condensates, both reconstituted in vitro and naturally in cells. These analyses will show whether condensates have organized internal structures, and if so, how these differ in natural and diseased conditions (e.g. in cells expressing PML-RAR fusions that cause promyelocytic leukemia, or mutants of the FUS protein that cause ALS). This work involves a combination of super-resolution fluorescence microscopy, as well as collaborative cryo-electron tomography. Ultimately our goal is to develop a comprehensive, quantitative understanding of the composition, dynamics and structure of representative biomolecular condensates, and to discover broadly applicable principles on how these structures function and evolve in biology.
For further details on specific projects we invite you to explore the descriptions found on individual lab members’ pages.