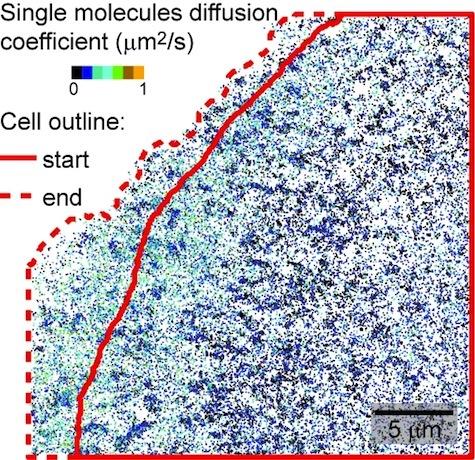
Inferring molecular trends and relationships from inherently heterogeneous data
A hallmark of single-cell and single-molecule data is the inherent heterogeneity in the measured properties. Therefore, our studies pose very exciting mathematical/statistical challenges:
- How to distinguish between true/meaningful biological variations, and fluctuations purely reflecting the inherent stochasticity of the system?
- How to distinguish between the inherent stochasticity of the system and noise due to measurement errors?
- How to expose underlying trends (in time and/or space) despite the multiple layers of stochasticity in the observed system (i.e. (1) and (2) just above)?
- How to extract relationships between various molecular and cellular properties, again given the multiple layers of stochasticity?
We are actively working on solving many of these challenges for our various ongoing projects, for example, characterizing molecular movement and interactions, or to derive cellular trends over time, or determining the relationship between molecular behavior and subcellular context, to name a few. For this, we are adapting and advancing approaches from statistics, machine learning, and even econometrics. Here is an example from our work to characterize molecular movement:
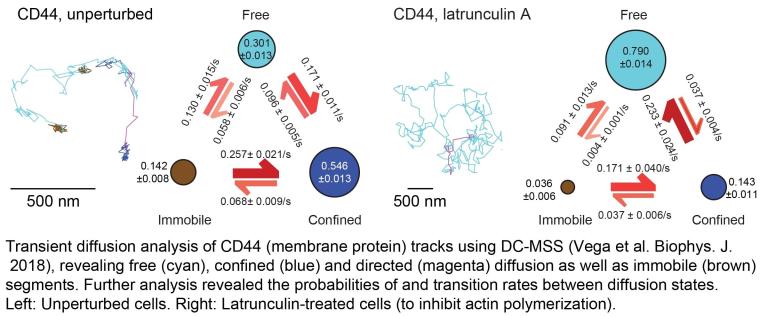
Linking molecular and cellular behaviors across spatial and temporal scales
Cellular behavior is the output of molecular behavior, yet linking the two to each other explicitly and quantitatively is challenging. This is because of their disparate spatial and temporal scales, and because cellular behavior is the emergent product of many molecules interacting with each other and many processes influencing each other. We are very fascinated by this challenge! In the past, we developed multi-scale analysis tools to link the dynamics of integrin molecules (key receptors in cell-matrix adhesion) to cell edge protrusion activity, in collaboration with the Galbraith Lab at Oregon Health and Science University (visual example shown in the image below).
In the context of our current projects, we are interested in the multi-scale analysis of receptor behavior and cellular signaling, receptor behavior, and actin cortex dynamics.