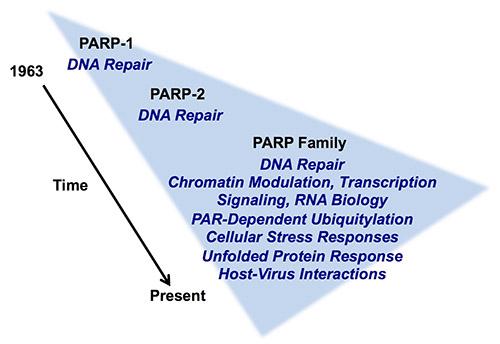
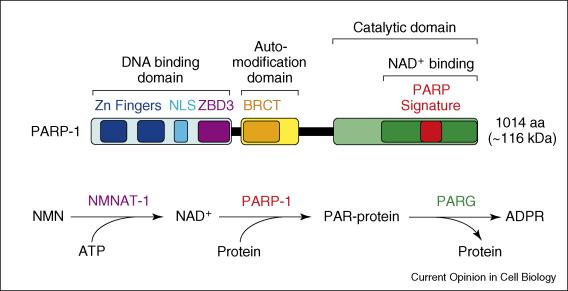
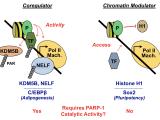
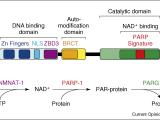
Studies of poly(ADP-ribosyl)ation by nuclear poly(ADP-ribose) polymerases (PARPs) enzymes were largely focused on their roles in DNA damage detection and repair in the 1960s through 1990s. In the early 2000s, the Kraus Lab was one of a small number that began to link PARP-1, an abundant nuclear PARP, to the regulation of chromatin structure and gene regulation. Using a range of biophysical, biochemical, and molecular approaches, the Kraus Lab found that PARP-1 plays key roles in the modulation of chromatin structure and gene expression in response to extracellular signals, such as those mediated by estrogens and TNFα. In addition, we have taken the lead in trying to understand how the synthesis of nuclear NAD+, the substrate for PARP-1, by the nuclear NAD+ synthase NMNAT-1 controls the gene regulatory functions of PARP-1 and downstream biological outcomes. These studies have linked cellular metabolic state, especially in the nucleus, to signal-regulated transcriptional outcomes.
Selected Publications
Krishnakumar R., Gamble M. J., Frizzell K. M., Berrocal J. G., Kininis M., Kraus W. L. (2008) Reciprocal binding of PARP-1 and histone H1 at promoters specifies transcriptional outcomes. Science 319:819-821. PMID: 18258916
Gamble M. J., Frizzell K. M., Yang C., Krishnakumar R., Kraus W. L. (2010) The histone variant macroH2A1 marks repressed autosomal chromatin, but protects a subset of its target genes from silencing. Genes & Development 24(1), 21-32. PMCID: PMC2802188
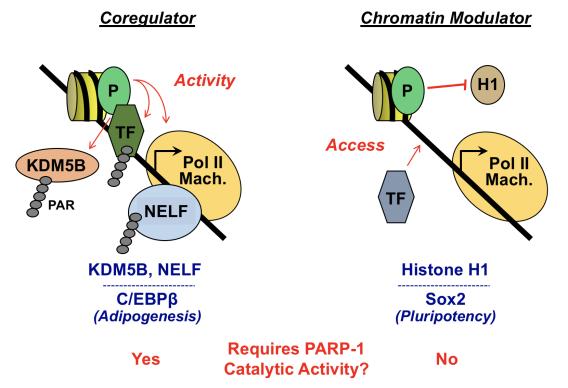
Krishnakumar R., Kraus W. L. (2010) PARP-1 Regulates chromatin structure and transcription through a KDM5B-dependent Pathway. Molecular Cell 39:736-749. PMCID: PMC2939044
Gibson B.A., Zhang Y., Jiang H., Hussey K.M., Shrimp J.H., Lin H., Schwede F., Yu Y., Kraus W.L. (2016) Chemical genetic discovery of PARP targets reveals a role for PARP-1 in transcription elongation. Science 353:45-50. PMID: 27256882
Liu Z., Kraus W.L. (2017) Catalytic-independent functions of PARP-1 determine Sox2 pioneer activity at intractable genomic loci. Molecular Cell 65: 589-603. PMID: 28212747
Luo X., Ryu K.W., Kim D.K., Nandu T., Medina C.J., Gupte R., Gibson B.A., Soccio R.E., Yu Y., Gupta R., Kraus W.L. (2017) PARP-1 controls the adipogenic transcription program by PARylating C/EPBβ and modulating its transcriptional activity. Molecular Cell 65:260-271. PMID: 28107648
Ryu K. W., Nandu T., Kim J., Challa S., DeBerardinis R. J., Kraus W. L. (2018) Metabolic regulation of transcription through compartmentalized NAD(+) biosynthesis. Science 360(6389):eaan5780. PMID: 29748257
Kim D.S., Camacho C.V., Nagari A., Malladi V.S., Challa S., Kraus W.L. (2019) Activation of PARP-1 by snoRNAs controls ribosome biogenesis and cell growth via the RNA helicase DDX21. Molecular Cell 75(6):1270-1285. PMID: 31351877
Huang D., Kim D.S., Kraus W.L. (2020) Specific binding of snoRNAs to PARP-1 promotes NAD+-dependent catalytic activation. Biochemistry 59(16):1559-1564. PMID: 32293172
Huang D., Camacho V.C., Setlem R., Ryu K.W., Parameswaran B., Gupta R.K., Kraus W.L. (2020) Functional Interplay between Histone H2B ADP-Ribosylation and Phosphorylation Controls Adipogenesis. Molecular Cell 79(6):934-939. PMID: 32822587
Palavalli Parsons L.H., Challa S., Gibson B.A., Nandu T., Stokes M.S., Huang D., Lea J.S., Kraus W.L. (2021) Identification of PARP-7 substrates reveals a role for MARylation in microtubule control in ovarian cancer cells. Elife 10:e60481. PMID: 33475085
Gupte R., Nandu T., Kraus W.L. (2021) Nuclear ADP-ribosylation drives IFNγ-dependent STAT1α enhancer formation in macrophages. Nature Communications 12(1):3931. PMID: 34168143
Challa S., Khulpateea B.R., Nandu T., Camacho C.V., Ryu K.W., Chen H., Peng Y., Lea J.S., Kraus W.L. (2021) Ribosome ADP-ribosylation inhibits translation and maintains proteostasis in cancers. Cell 184:4531-4546.e26. PMCID: PMC8380725
Huang D., Kraus W.L. (2022) The expanding universe of PARP1-mediated molecular and therapeutic mechanisms. Molecular Cell 82(12):2315-2334. PMID: 35271815
Huang D., Camacho C.V., Martire S., Nagari A., Setlem R., Gong X., Edwards A.D., Chiu S.P., Banaszynski L.A., Kraus W.L. (2022) Oncohistone Mutations Occur at Functional Sites of Regulatory ADP-Ribosylation. Cancer Research 82(13):2361-2377. PMID 35472077
Challa S., Ryu K.W., Whitaker A.L., Abshier J.C., Camacho C.V., Kraus W.L. (2022) Development and characterization of new tools for detecting poly(ADP-ribose) in vitro and in vivo. Elife 11:e72464. PMID: 35476036
Jones A., Kraus W.L. (2022) Multiomics analysis of the NAD+-PARP1 axis reveals a role for site-specific ADP-ribosylation in splicing in embryonic stem cells. Cell Reports 36(9-10):601-617. PMID 35654456